Post-transcriptional modification
This is missing information about sections for RNA other than mRNA. (October 2020) |
| Part of a series on |
| Genetics |
|---|
 |
Transcriptional modification or co-transcriptional modification is a set of biological processes common to most eukaryotic cells by which an RNA primary transcript is chemically altered following transcription from a gene to produce a mature, functional RNA molecule that can then leave the nucleus and perform any of a variety of different functions in the cell.[1] There are many types of post-transcriptional modifications achieved through a diverse class of molecular mechanisms.
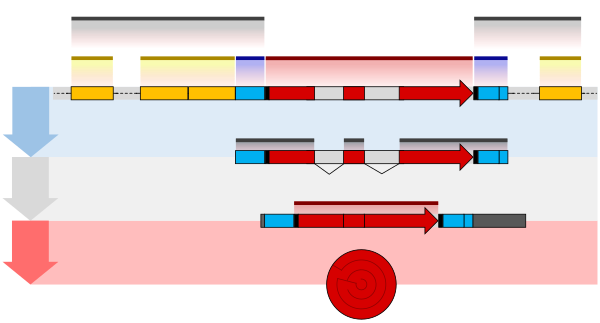
One example is the conversion of precursor messenger RNA transcripts into mature messenger RNA that is subsequently capable of being translated into protein. This process includes three major steps that significantly modify the chemical structure of the RNA molecule: the addition of a 5' cap, the addition of a 3' polyadenylated tail, and RNA splicing. Such processing is vital for the correct translation of eukaryotic genomes because the initial precursor mRNA produced by transcription often contains both exons (coding sequences) and introns (non-coding sequences); splicing removes the introns and links the exons directly, while the cap and tail facilitate the transport of the mRNA to a ribosome and protect it from molecular degradation.[2]
Post-transcriptional modifications may also occur during the processing of other transcripts which ultimately become transfer RNA, ribosomal RNA, or any of the other types of RNA used by the cell.
mRNA processing
[edit]|
|
5' processing
[edit]Capping
[edit]Capping of the pre-mRNA involves the addition of 7-methylguanosine (m7G) to the 5' end. To achieve this, the terminal 5' phosphate requires removal, which is done with the aid of enzyme RNA triphosphatase. The enzyme guanosyl transferase then catalyses the reaction, which produces the diphosphate 5' end. The diphosphate 5' end then attacks the alpha phosphorus atom of a GTP molecule in order to add the guanine residue in a 5'5' triphosphate link. The enzyme (guanine-N7-)-methyltransferase ("cap MTase") transfers a methyl group from S-adenosyl methionine to the guanine ring.[4] This type of cap, with just the (m7G) in position is called a cap 0 structure. The ribose of the adjacent nucleotide may also be methylated to give a cap 1. Methylation of nucleotides downstream of the RNA molecule produce cap 2, cap 3 structures and so on. In these cases the methyl groups are added to the 2' OH groups of the ribose sugar. The cap protects the 5' end of the primary RNA transcript from attack by ribonucleases that have specificity to the 3'5' phosphodiester bonds.[5]
3' processing
[edit]Cleavage and polyadenylation
[edit]The pre-mRNA processing at the 3' end of the RNA molecule involves cleavage of its 3' end and then the addition of about 250 adenine residues to form a poly(A) tail. The cleavage and adenylation reactions occur primarily if a polyadenylation signal sequence (5'- AAUAAA-3') is located near the 3' end of the pre-mRNA molecule, which is followed by another sequence, which is usually (5'-CA-3') and is the site of cleavage. A GU-rich sequence is also usually present further downstream on the pre-mRNA molecule. More recently, it has been demonstrated that alternate signal sequences such as UGUA upstream off the cleavage site can also direct cleavage and polyadenylation in the absence of the AAUAAA signal. These two signals are not mutually independent, and often coexist. After the synthesis of the sequence elements, several multi-subunit proteins are transferred to the RNA molecule. The transfer of these sequence specific binding proteins cleavage and polyadenylation specificity factor (CPSF), Cleavage Factor I (CF I) and cleavage stimulation factor (CStF) occurs from RNA Polymerase II. The three factors bind to the sequence elements. The AAUAAA signal is directly bound by CPSF. For UGUA dependent processing sites, binding of the multi protein complex is done by Cleavage Factor I (CF I). The resultant protein complex formed contains additional cleavage factors and the enzyme Polyadenylate Polymerase (PAP). This complex cleaves the RNA between the polyadenylation sequence and the GU-rich sequence at the cleavage site marked by the (5'-CA-3') sequences. Poly(A) polymerase then adds about 200 adenine units to the new 3' end of the RNA molecule using ATP as a precursor. As the poly(A) tail is synthesized, it binds multiple copies of poly(A)-binding protein, which protects the 3'end from ribonuclease digestion by enzymes including the CCR4-Not complex.[5]
Intron splicing
[edit]RNA splicing is the process by which introns, regions of RNA that do not code for proteins, are removed from the pre-mRNA and the remaining exons connected to re-form a single continuous molecule. Exons are sections of mRNA which become "expressed" or translated into a protein. They are the coding portions of a mRNA molecule.[6] Although most RNA splicing occurs after the complete synthesis and end-capping of the pre-mRNA, transcripts with many exons can be spliced co-transcriptionally.[7] The splicing reaction is catalyzed by a large protein complex called the spliceosome assembled from proteins and small nuclear RNA molecules that recognize splice sites in the pre-mRNA sequence. Many pre-mRNAs, including those encoding antibodies, can be spliced in multiple ways to produce different mature mRNAs that encode different protein sequences. This process is known as alternative splicing, and allows production of a large variety of proteins from a limited amount of DNA.
Histone mRNA processing
[edit]Histones H2A, H2B, H3 and H4 form the core of a nucleosome and thus are called core histones. Processing of core histones is done differently because typical histone mRNA lacks several features of other eukaryotic mRNAs, such as poly(A) tail and introns. Thus, such mRNAs do not undergo splicing and their 3' processing is done independent of most cleavage and polyadenylation factors. Core histone mRNAs have a special stem-loop structure at 3-prime end that is recognized by a stem–loop binding protein and a downstream sequence, called histone downstream element (HDE) that recruits U7 snRNA. Cleavage and polyadenylation specificity factor 73 cuts mRNA between stem-loop and HDE[8]
Histone variants, such as H2A.Z or H3.3, however, have introns and are processed as normal mRNAs including splicing and polyadenylation.[8]
See also
[edit]References
[edit]- ^ Kiss T (July 2001). "Small nucleolar RNA-guided post-transcriptional modification of cellular RNAs". The EMBO Journal. 20 (14): 3617–22. doi:10.1093/emboj/20.14.3617. PMC 125535. PMID 11447102.
- ^ Berg, Tymoczko & Stryer 2007, p. 836
- ^ a b Shafee, Thomas; Lowe, Rohan (2017). "Eukaryotic and prokaryotic gene structure". WikiJournal of Medicine. 4 (1). doi:10.15347/wjm/2017.002. ISSN 2002-4436.
- ^ Yamada-Okabe T, Mio T, Kashima Y, Matsui M, Arisawa M, Yamada-Okabe H (November 1999). "The Candida albicans gene for mRNA 5-cap methyltransferase: identification of additional residues essential for catalysis". Microbiology. 145 ( Pt 11) (11): 3023–33. doi:10.1099/00221287-145-11-3023. PMID 10589710.
- ^ a b Hames & Hooper 2006, p. 221
- ^ Biology. Mgraw hill education. 2014. pp. 241–242. ISBN 978-981-4581-85-1.
- ^ Lodish HF, Berk A, Kaiser C, Krieger M, Scott MP, Bretscher A, Ploegh H, Matsudaira PT (2007). "Chapter 8: Post-transcriptional Gene Control". Molecular Cell .Biology. San Francisco: WH Freeman. ISBN 978-0-7167-7601-7.
- ^ a b Marzluff WF, Wagner EJ, Duronio RJ (November 2008). "Metabolism and regulation of canonical histone mRNAs: life without a poly(A) tail". Nature Reviews. Genetics. 9 (11): 843–54. doi:10.1038/nrg2438. PMC 2715827. PMID 18927579.
Further reading
[edit]- Berg JM, Tymoczko JL, Stryer L (2007). Biochemistry (6 ed.). New York: WH Freeman & Co. ISBN 978-0-7167-6766-4.
- Hames D, Hooper N (2006). Instant Notes Biochemistry. Vol. 58 (3 ed.). Leeds: Taylor and Francis. p. 767. ISBN 978-0-415-36778-3. PMID 11098183.
{{cite book}}:|journal=ignored (help) - Sun WJ, Li JH, Liu S, Wu J, Zhou H, Qu LH, Yang JH (January 2016). "RMBase: a resource for decoding the landscape of RNA modifications from high-throughput sequencing data". Nucleic Acids Research. 44 (D1): D259-65. doi:10.1093/nar/gkv1036. PMC 4702777. PMID 26464443.
- Machnicka MA, Milanowska K, Osman Oglou O, Purta E, Kurkowska M, Olchowik A, Januszewski W, Kalinowski S, Dunin-Horkawicz S, Rother KM, Helm M, Bujnicki JM, Grosjean H (January 2013). "MODOMICS: a database of RNA modification pathways--2013 update". Nucleic Acids Research. 41 (Database issue): D262-7. doi:10.1093/nar/gks1007. PMC 3531130. PMID 23118484.
- Cantara WA, Crain PF, Rozenski J, McCloskey JA, Harris KA, Zhang X, Vendeix FA, Fabris D, Agris PF (January 2011). "The RNA Modification Database, RNAMDB: 2011 update". Nucleic Acids Research. 39 (Database issue): D195-201. doi:10.1093/nar/gkq1028. PMC 3013656. PMID 21071406.
- Post-Transcriptional+RNA+Modification at the U.S. National Library of Medicine Medical Subject Headings (MeSH)